Maize
molecular
Atlas
Navigating genetic diversity
Understanding and using diversity
The molecular atlas provides data, tools and resources to allow germplasm bank users; maize breeders, researchers, extension agents etc., to identify the diversity of likely value for their specific needs from the tens of thousands of landraces available to them.
The atlas is structured around three pillars; data and knowledge, tools and training resources. The specific components within these pillars can be accessed alone or in combination, as required by users as illustrated in the example of identification of new variation for heat tolerance.
We are interested in receiving your feedback about the molecular atlas. Please feel free to contact us to ask questions or provide comments.
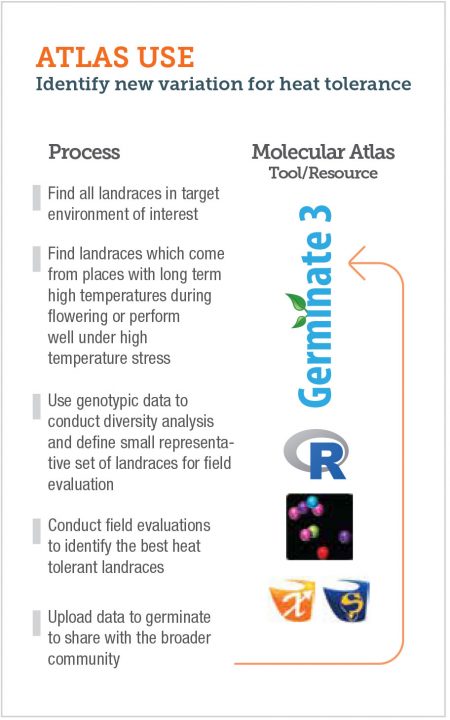
Data, Knowledge & training resources

SeeD Maize Germinate is a genetic resources database that stores and provides search and visualization tools for passport data, accession collection information and data, genotypic data and phenotypic data for maize accessions and derived materials studied as part of the initiative. Germinate was developed by James Hutton Institute, in collaboration with CIMMYT.
Software tools


CurlyWhirly is a multi-platform application developed to enable interactive visualization of multi-variate analysis, with a particular focus on the outputs of Principal Coordinate Analysis, Principal Components Analysis, Canonical analysis and Multidimensional Scaling. Integration of results with other categorical data facilitates interpretation and selection of points of interest. CurlyWhirly was developed by James Hutton Institute, in collaboration with CIMMYT.
TRAINING RESOURCES
Links to other maize resources
We are well aware that SeeD is only one of several projects and online resources that deal with maize genetic diversity or molecular data. As we develop the Maize Molecular Atlas, we will attempt to link it to other on-line resources and projects, such as:
- The AMAIZING project
- The MaizeGDB portal
- The Panzea website
- The Maize Genome Database
- The data generated by the Latin American Maize Project (LAMP)
- The data generated by the Global Maize Landrace Project
- The Master Project for Mexican Maize (PMMM)

















